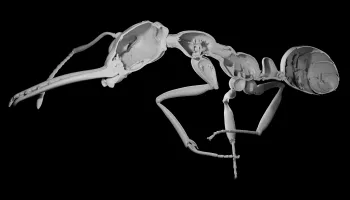
Researchers leveraged advanced technologies and artificial intelligence to hasten the process of generating 2,000 3D digital ant images. Now, a class of UMD computer science majors is working to bring the data to life.
For more than a decade, Evan Economo’s lab has been using micro-CT machines to scan insect specimens. The resulting X-ray images help researchers study the form and structure of insects—a subfield of entomology known as morphology—but the process is costly and time-consuming.

“One limitation is that you can get this rich 3D dataset, but it could take 10 hours to scan one specimen,” explained Economo, who chairs the University of Maryland’s Department of Entomology and holds the James B. Gahan and Margaret H. Gahan Professorship.
As a senior author of a paper published in the journal Nature Methods on March 5, 2026, Economo tested a high-tech workflow to speed up their efforts. A research team co-led by Economo and Thomas van de Kamp at the Karlsruhe Institute of Technology (KIT) in Germany combined a Synchrotron particle accelerator, X-ray scanning, robotics, and artificial intelligence (AI) to create interactive digital images representing 800 different ant species.
Ultimately, these technologies enabled the team to drastically reduce the time it takes to scan specimens and transform raw image files into high-resolution 3D models.
“We’ve estimated that if we were to carry out this project with a lab-based CT scanner, it would take six years of continuous operation,” said Julian Katzke, the study’s first author and graduate of Economo’s lab at the Okinawa Institute of Science and Technology (OIST) in Japan. “With the setup at KIT, we scanned 2,000 specimens in a single week.”
Dubbed Antscan, this project could serve as a blueprint for future digitization efforts—not just for ants, but for a wide variety of species. The raw files for constructing 3D models are free for anyone to download, and a built-in viewer of every ant allows for easy access to the finished 3D images.
“The value of this study is not only about ants—it's much broader,” said Economo, who is now an adjunct professor at OIST in addition to his UMD role. “When specimens are digitized, we can build libraries of organisms that can streamline their use from scientific laboratories to classrooms to Hollywood studios.”
‘A living library’
To build such an expansive digital library, the research team sourced ethanol-preserved ant specimens from partner institutions, museum collections and experts around the world. After the researchers sorted the specimens by species and caste, they brought them to KIT for high-throughput X-ray micro-CT scanning, which is comparable to medical CT scans but in much higher magnification.

A synchrotron particle accelerator produced a high-intensity X-ray beam to rapidly scan a huge number of specimens, and a robotic sample changer rotated and swapped out the specimens every 30 seconds. This enabled the creation of 2D image stacks that could then be used to construct 3D models.
While useful, the raw image files depicted ant specimens in contorted poses—a far cry from the lifelike models that researchers hoped to build. As a follow-up to Antscan, students in UMD Computer Science Associate Professor James Purtilo’s CMSC 435: “Software Engineering” course are using AI to automate the process of “pose estimation,” enabling awkward ant poses to be transformed into natural ones that might be seen in the wild.
“This collaboration was a great opportunity for us,” Purtilo said. “A capstone is intended to challenge students to integrate skills, function as an effective team and demonstrate their ability to solve real problems. And this problem was a doozy.”
Antscan’s 3D images reveal internal structures like muscles, nervous systems, digestive systems and stingers at micrometer resolution. The models can easily be animated or incorporated into virtual reality worlds for research, education or entertainment.
“To do this manually would have taken years, so without these computational tools it basically would never have been done,” Economo said. “Now, we are making large strides toward creating a living library of interactive models corresponding to Earth’s biodiversity. AI will enable us to explore the diversity of life and share it with the world.”
Antscan in action
The database has already begun to demonstrate its scientific value. Economo was the senior author of a paper published in the journal Science Advances on December 19, 2025, in which researchers used Antscan data to explore whether ant colonies would fare better with a higher number of weaker workers or fewer, more robust workers.

Economo and collaborators studied the correlation between cuticle volume, colony size and colony diversification across more than 500 ant species. The cuticle, which forms the protective outer layer of the exoskeleton, is nutritionally expensive in nitrogen and various minerals, meaning that thicker armor requires a greater investment of resources per ant.
The team found a strong negative correlation between cuticle volume and colony size, suggesting that prioritizing the quantity of ants over the quality of their armor facilitates the development of larger and more diverse ant societies.
In this case, Antscan enabled researchers to take a closer look at cuticle volume, which has been difficult to measure prior to these models. The Antscan project also covered the same ant species as a June 2025 study in the journal Cell, co-authored by Economo, that produced a set of high-quality ant genomes. Combined, these studies could enable more complex research into the relationship between morphological data and genomic variation.
Given their high fidelity, the scans could someday even be used to train machine learning models to accurately detect ants in the field for observational studies of their behavior. Going forward, Economo plans to scan more specimens into the system while continuing to work with UMD computer science students to apply these AI techniques to new datasets.
“This work moves us further into the big data era of capturing, analyzing and sharing organismal shape and form,” Economo said. “The potential for integrating these data with other data types and technologies is immense and very exciting.”
###
Their paper, “High-throughput phenomics of global ant biodiversity,” was published in the journal Nature Methods on March 5, 2026.
This article was adapted from text provided by the Okinawa Institute of Science and Technology.
This research was supported by the German Ministry for Research and Education; the Ministry of Science, Research and the Arts Baden-Württemberg; the German Research Foundation (Grant Nos. INST 35/1503-1 FUGG and 502787686); the Okinawa Institute of Science and Technology Graduate University; the Japan Society for the Promotion of Science (Grant Nos. 18K14768, 21K06326 and 22KJ3077); the Australian Research Council (Award No. IC 180100008); HUN-REN Hungarian Research and the National Research, Development, and Innovation Fund (Grant No. K 147781); the Conselho Nacional de Desenvolvimento Científico e Tecnológico (Grant No. 301495/2019-0); the Critical Ecosystem Partnership Fund, a joint initiative of l’Agence Française de Développement, Conservation International, the European Union, the Global Environment Facility, the government of Japan and the World Bank; the U.S. National Science Foundation (Grant Nos. DEB-1932467, DEB 1927161 and IOS-2128304); the Italian Ministry for University and Research; the Environment and Conservation Fund in Hong Kong (Award No. Nb. ECF 137/2020); and Fundação para a Ciência e a Tecnologia. This article does not necessarily reflect the views of these organizations.