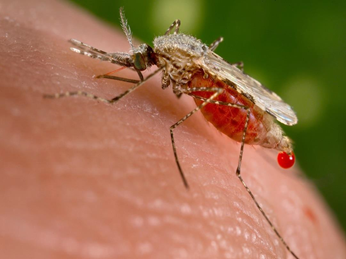
 Figure 1. Anopheles, the mosquito that transmits human malaria (photo from Wikipedia) Figure 1. Anopheles, the mosquito that transmits human malaria (photo from Wikipedia) Researchers are developing new genomic approaches to detect pathogens and their vectors. These new methods reduce cost, increase sensitivity, and may allow for early detection of new parasites. 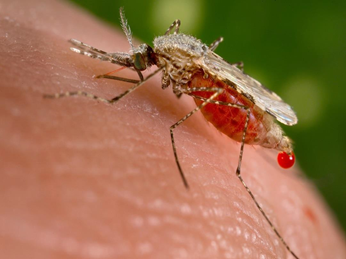 Figure 1. Anopheles, the mosquito that transmits human malaria (photo from Wikipedia) Figure 1. Anopheles, the mosquito that transmits human malaria (photo from Wikipedia) Insect-transmitted diseases are a major health concern globally. One of the most widely studied insects responsible for numerous deaths each year due to the transmission of eukaryotic parasites is Anopheles. Did you know that Anopheles is capable of transmitting malaria? Anopheles is the vector of human malaria, and is found on all continents with over 500 species worldwide. Malaria is a debilitating disease induced by parasites which are transmitted from person to person through the infectious bites of female Anopheles mosquitoes. For many years, researchers have been searching for a way to minimize disease transmission and stopping outbreaks, but were faced with numerous roadblocks. One of these researchers, Dr. David Serre, an Associate Professor at the University of Maryland School of Medicine has been working diligently in his lab to come up with new genomics approaches to better survey mosquito feeding behavior, species diversity, and eukaryotic parasites. Without blood feeding, female mosquitoes cannot attain the nutrients essential for egg production. Blood feeding is also necessary for parasites to move from one host to another and complete their life cycle. Thus, understanding the precise feeding behavior of mosquitoes is critical for disrupting this process. To understand the vector feeding behavior by Anopheles, Dr. Serre developed a method of using Next-Generation Sequencing (NGS) to identify what species mosquitoes fed upon by sequencing DNA in the blood meals they had taken. All mammalian 16 S rRNAs in the mixed blood meals from multiple individual mosquitoes were amplified by Polymerase Chain Reaction (PCR) using universal primers designed to target more than 95% of the mammalian 16S rRNA sequences present in the NCBI nucleotide database. PCR products from up to 96 mosquitoes were pooled and sequenced simultaneously with only about 7-9 hours of hands-on time required, which provides a comprehensive and quantitative perspective on the composition of each blood meal. Interestingly, more than one species DNA were found in one mosquito. This showed a much wider variety of hosts than previously recorded and opened new insights into mosquito feeding behavior. Anopheles species have become increasingly prevalent on all continents, though they remain poorly described. Factors such as mosquito size, species diversity, and near identical appearance between species may account for the lack of comprehensive studies at the species level. To address these issues, whole mitochondrial DNA (mtDNA) of Anopheles mosquitoes was extracted and amplified by long-range PCR to study the genetic diversity of mosquitoes collected. This novel use of NGS combined with their preliminary findings can guide efforts in detection of reservoirs of vector-borne disease and improving methods of deterring or eliminating pathogen transmission. Current techniques are costly in labor and are highly specific to individual parasites. Dr. Serre’s aim is to not only reduce cost and expand detection to a majority of parasites, but to do so in one test, which will also allow for early detection of emerging pathogens. Having laid the ground work for understanding feeding behavior and mosquito diversity, the remaining piece of the puzzle involves the reason they’re interested: the parasites. Dr. Serre’s group has utilized a high-throughput screening approach for detection of DNA from eukaryotic parasites as well. High-throughput screening is a recent technique where numerous samples are tested at once under the same experimental conditions, and it allows them to meet their goal of reducing cost and widening detection parameters. Serre’s group has collected tens of thousands of mosquitoes from over 25 species using hundreds of CO2-baited light traps in Maryland. His group first isolated DNA from all mosquitoes collected in a given trap. Then, they searched this DNA for the presence of eukaryotic parasites by PCR amplification followed by Illumina high-throughput sequencing methods. This allows for screening of all eukaryotic parasites with known genetic sequences, such as nematodes and microsporidians, captured in every trap. They again experienced the same benefits with their previous studies on mosquito feeding behavior and diversity, successfully catching all known parasites with their detection methods.
The significance of this work is that they are laying the groundwork for collecting and testing massive amounts of mosquitoes for feeding behavior, diversity, and distinct parasites. Starting right here in Maryland, they are beginning to build a database of mosquito species and parasite genomes while refining the sensitivity of their genetic techniques. Ultimately, the information from the database can be used to guide the description of previously unknown eukaryotic parasites in mosquitoes, as well as understand their frequency and spatial distribution. Surveillance of insects, especially those that host pathogens, is pivotal in alleviating the burden of diseases transmitted by insects and stopping outbreaks from occurring. Current surveillance methods in use are quite costly in resources and only target specific pathogens carried by the most threatening mosquito species. Dr. Serre’s work can be used to develop new surveillance methods that harness modern sequencing technology. This could prove useful in identifying and describing unknown and known eukaryotic parasites and their insect hosts. Dr. Serre’s methods would also allow for faster, more cost-effective, and detailed surveillance of insect-transmitted diseases throughout the world. Future work under this approach includes expansion to other insect species like honeybees, and may expand to the detection of other pathogens such as viruses, bacteria, fungi, and nematodes as well. This further displays the potential impact of these new methods. By developing a standardized method, which is fast and inexpensive, for vector-borne pathogens, Dr. Serre and his group put humanity one step closer to not only eradicating many human diseases but also diseases among insects and other animals. Bloggers: Mike Nan is a PhD student in the St. Leger Lab. He is interested in understanding the genetics and molecular biology of entomopathogenic fungi using genetic engineering and genomics. Anthony Nearman is a PhD student in the vanEngelsdorp Lab. He studies honey bee nutrition and immunological responses to invading pathogens. References
Comments are closed.
|
Categories
All
Archives
June 2024
|
Department of Entomology
University of Maryland
4112 Plant Sciences Building
College Park, MD 20742-4454
USA
Telephone: 301.405.3911
Fax: 301.314.9290
University of Maryland
4112 Plant Sciences Building
College Park, MD 20742-4454
USA
Telephone: 301.405.3911
Fax: 301.314.9290

 RSS Feed
RSS Feed




